Whole Genome Sequencing/ Whole Exome Sequencing Working Group Update
April 15, 2021
Written by Guillaume Butler-Laporte, Minttu Marttila, Kumar Veerapen
Note: This update is geared towards the scientific community as an update to note the sequencing analysis efforts in COVID-19 host genetics. If you have further enquiries, please send us an email at hgi-faq@icda.bio —we’d be happy to update the information here to provide more clarity.
The COVID-19 Host Genetics Initiative’s (COVID-19 HGI) has focused on common genetic determinants of COVID-19. However, the role of rare genetic determinants (found in less than 0.5% of the population) remains unclear to the etiology of COVID-19 infections. Based on our knowledge of complex diseases, some rare variants can be associated with extreme clinical presentations. In the context of COVID-19, the characterization of rare variants may provide great insights into explaining the biological mechanisms of patients with severe COVID-19.
In order to study these variants, we need a different type of technology to “read” a patients’ genetic code. This technology is usually referred to as “sequencing” (as opposed to the “genotyping” technology that is currently being used in the COVID-19 HGI). Sequencing allows us to determine genetic variants carried by every patient with improved precision. For the extension that this blog post focuses on, we are employing sequencing of the entire genome (whole-genome-sequencing, WGS) or restricted to the protein coding segments of the genome (whole-exome-sequencing, WES).
Challenges in rare genetic variant analysis
Studying rare variants is difficult: only a few (if any) patients in each cohort will carry a rare variant that has a high likelihood of truly causing severe COVID-19. This leads to the following challenges:
- Large sample sizes are required to identify rare pathogenic variants.
- The available genome-wide association study (GWAS) statistical methods perform poorly when dealing with rare events.
- Finding rare variants with meaningful biological etiology and pooling patients’ rare variants across multiple cohorts poses a logistical challenge.
COVID-19 Host Genetics and Rare Variants: The WES/WGS Working Group
Therefore, the WES/WGS working group is divided into two main tracks aiming to use data from a large collaborative of participating studies in order to improve statistical power of detecting meaningful rare genetic determinants in COVID-19 infections:
The first track uses gene “burden” tests to identify genes associated with COVID-19. In burden testing, instead of looking for the association of genetic variants to diseases/traits, we instead “pool” all pathogenic rare variants in each gene for each patient, and measure if patients with an enrichment of these variants within each gene are overrepresented in the disease trait or not. This alleviates the problem of decreased statistical method performance if we were to use usual single variant association tests done in GWASs. To further address this problem, burden testing will be done using Firth regression, which provides unbiased and stable association tests even with very small sample sizes.
The burden test used in the analysis is similar to an additive effect model used in GWAS, where each patient is given a score of 0, 1, or 2 at each gene depending on whether or not the gene contains a potentially pathologic variant (and if it is homo- or hetero-zygous). This allows us to use the same meta-analysis methods as in the COVID-19 HGI GWAS, allowing each cohort to perform their analyses locally, and bypassing many logistic problems related to data sharing.
As we plan to use this burden testing in severe COVID-19, we will therefore be improving our chances of identifying genes involved in severe COVID-19. Moreover, this analysis is fairly frequently used in the field of rare genetic analysis and can therefore be quickly performed by each of the study cohort analytical teams using Hail and REGENIE. This project is led by Brent Richards and Guillaume Butler-Laporte from McGill University.
The second track uses a federated learning approach to centrally coordinate analyses across cohorts, and leverage our knowledge of common variants to identify genes and rare variants involved in COVID-19. This effort is led by Alessandra Renieri and Francesca Mari from Siena University.
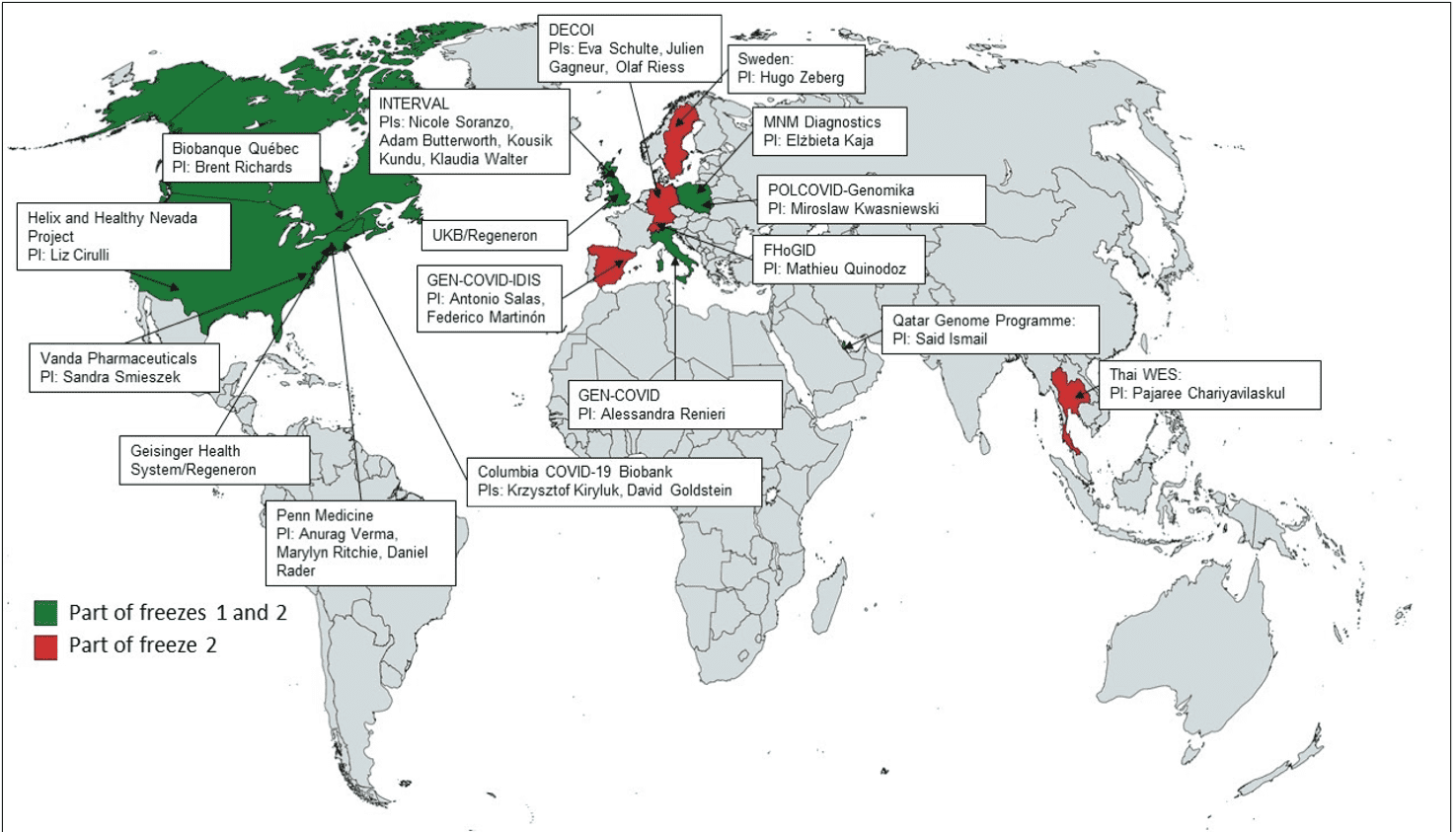
The WES/WGS working group had our first data freeze on December 17th 2020. Thus far, 7 studies have contributed: Gen-Covid, Columbia Biobank, Helix, Pol-Covid, Regeneron (UK Biobank and Geisinger Health System), Qatar Genome, Penn Medicine, and the Biobanque Qébécoise de la COVID-19 (Figure 1).
Figure 1: Map of projects and project leaders taking part in Whole Genome Sequencing/ Whole Exome Sequencing Working Group Data Freeze 2 (as of March 7, 2021). In green: countries who were represented in Data Freeze 1. In red: new countries for Data Freeze 2.
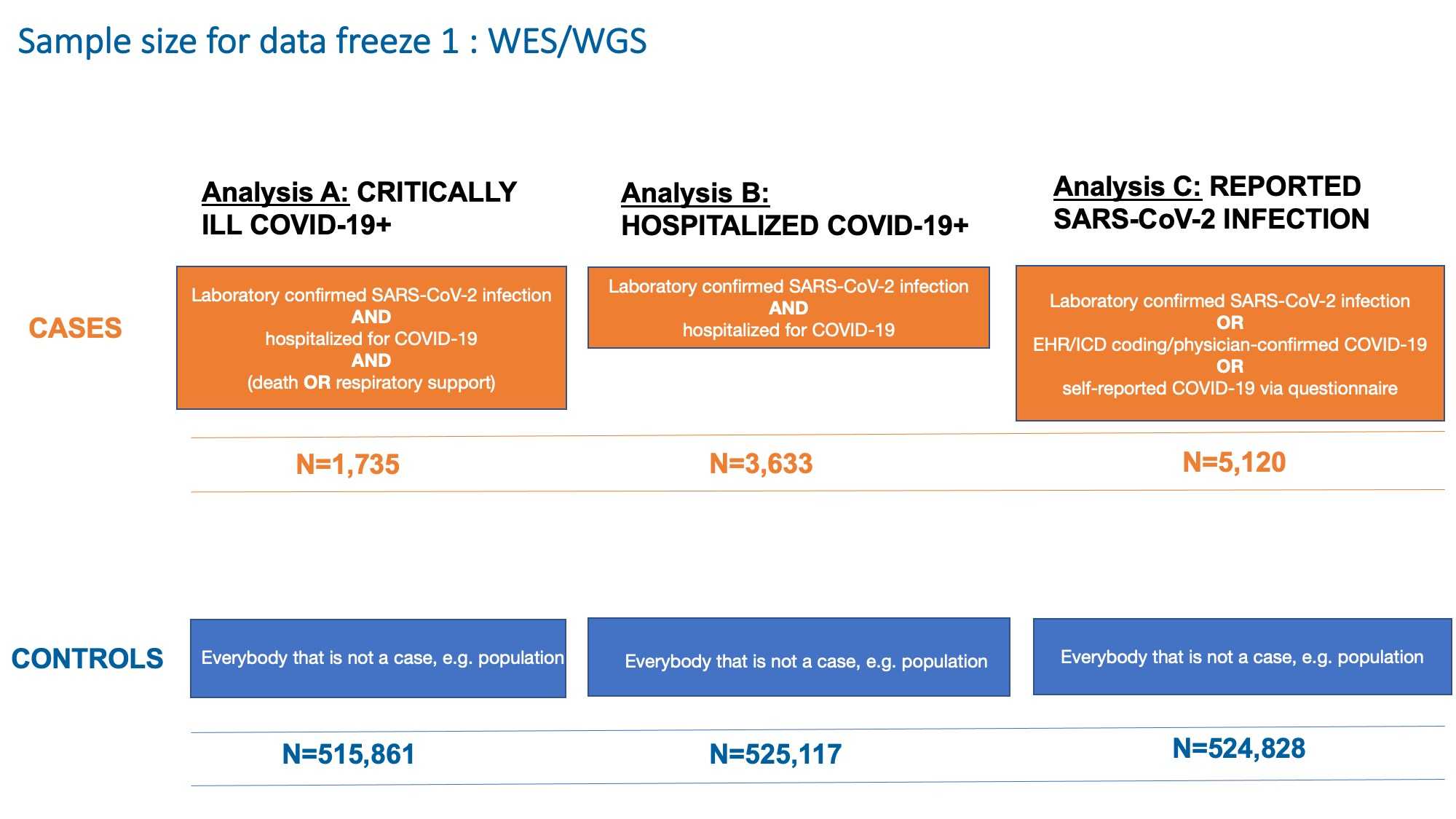
In this data freeze, we had a total of more than 0.5 million individuals for our analyses across analysis A, B, and C phenotype definitions (Figure 2).
Figure 2: Case-control distribution used in the sequencing efforts by the COVID-19 HGI
The analysis in this data freeze was restricted to the loci found in the COVID-19 HGI GWAS. However, we did not produce robust genetic associations. Despite our best efforts at applying stringent quality control and analytical protocols in our analysis, we hypothesize that this may be caused by a lack of statistical power in detecting significant genetic signals.
Improving Detection of Robust Genetic Signals in Data Freeze 2
Therefore, to circumvent our lack of sample size, we have instituted a second data freeze on March 1, 2021. As of the publication of this blog post, we have increased the number of studies from 7 to 16, and the number of represented countries from 5 to 11.
On top of improving our sample size, we are analyzing our data across all the protein coding regions of the genome i.e. the exome.
If you would like to collaborate, please contact the working group through e-mail: brent.richards@mcgill.ca or guillaume.butler-laporte@mail.mcgill.ca and on the COVID-19 HGI Slack channel: #covid-19-hg-wes-wgs.